A little bit of history…
The probability theory that is the foundation of Bayesian statistics was developed by Thomas Bayes (1702-1761). His ideas were accepted by some and challenged by others, and it was not until the mid-20th century that these ideas started gaining popularity. However, Bayesian methods were not widely implemented until about 1990, in part due to the complexity of the mathematical calculations. As computer power increased and new computing algorithms were developed, Bayesian statistics became more accessible; the models were not limited to simple models as in years past. Nowadays, we can fit complicated Bayesian models to almost every situation.Overview
Unlike in frequentist or classical statistics, in Bayesian analysis we treat the data as non-random observed quantities and the parameters as random. In classical statistics, the parameter is fixed, the data are a random sample, and we compute point estimates and confidence intervals for the parameters. On the other hand, in Bayesian statistics we compute a posterior density distribution, which represents the uncertainty in that parameter; i.e., what values the parameter is likely to take on. This probability distribution relies not only on the observed data, but also on the prior information at hand. This prior knowledge can be derived from previous data, similar systems, expert knowledge/opinion, or “gut instinct.” Therefore, the posterior distribution characterizes what we know and incorporates new evidence about the parameter after the data are observed. Using Bayes’ rule, the posterior distribution \((p(\theta | data))\) is computed as: [latex] p(\theta | data) \propto p(data | \theta) p(\theta) [/latex] where \(p(θ)\) is the prior distribution of the parameter \(θ\), \(p(data|θ)\) is the likelihood distribution, and the symbol \(\propto\) can be read as “is proportional to.” The prior distribution represents the previous knowledge of or belief regarding the parameter before the data are collected, whereas the likelihood is the probability of the observed data under certain model. In general, Bayesian analysis can be summarized in three steps:- Specify the prior and likelihood distributions
- Compute the posterior distribution, \(p(θ|data)\)
- Perform model diagnostics
Prior and Likelihood Distributions
Remember that the likelihood associated with a model is the probability of the observed data under such model. In Bayesian statistics, we combine that likelihood with the prior information in order to compute a posterior distribution. If we are confident in the previous information at hand, we could add it to the model by using an informative prior distribution. On the other hand, if we do not know much about the parameter, we could use a vague or diffuse prior distribution. Figure 1 shows two posterior distributions, reflecting the observed 6 successes we observed in 24 trials, and using a Beta distribution as a prior (see Beta-Binomial). The posterior distribution at the top of the figure was computed using vague prior distribution, and the posterior distribution at the bottom was obtained using an informative prior. Notice how the posterior variability decreases and the credible intervals (dashed vertical lines) get tighter when we use an informative prior over a vague one.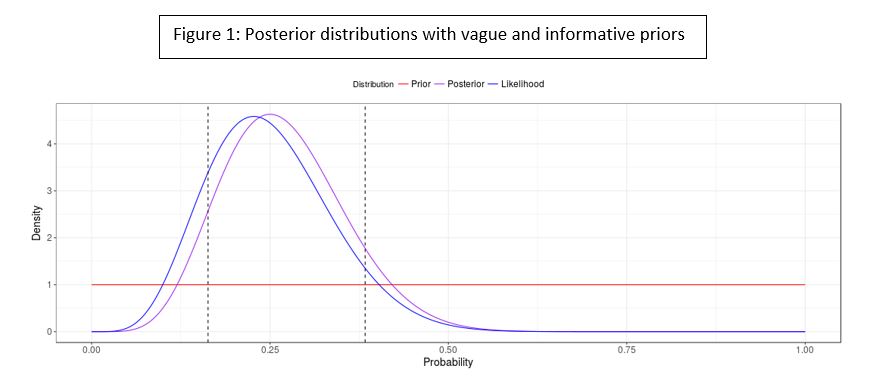
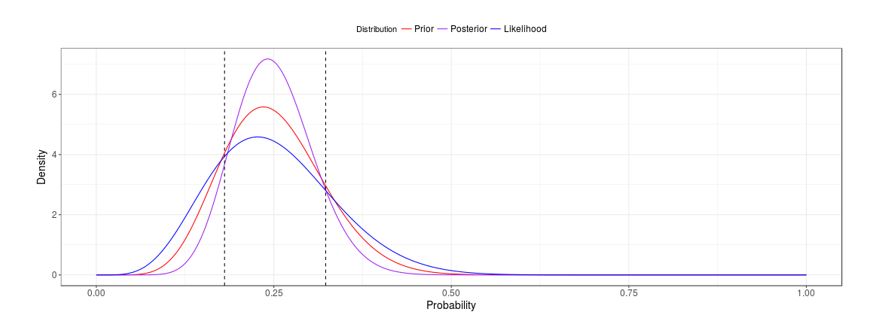 Regardless of the amount of information we have or how confident we are in that information, the prior distribution should at least take into account the domain (set of possible values) of the parameter. For example, if we want to estimate a proportion, the prior distribution for the parameter should range between 0 and 1.
Prior distributions could also be classified as conjugate priors. A conjugate prior is a prior that, when combined with the likelihood, results in a posterior distribution of the same form (parametric family) as the prior. An example of a conjugate prior is the beta prior for a binomial likelihood; the posterior distribution is also a beta distribution (see beta-binomial and Conjugate Prior). Conjugate priors are useful in the sense that the posterior distribution is of a known parametric form, thus the computations are less complicated and they can be interpreted as additional data. Even though conjugate priors have their advantages, their use is limited in real-world problems because they may not represent the prior knowledge appropriately for a specific situation, or no conjugate family exists for the real-world model. In situations where the form of the posterior distribution is not known (non-conjugacy), we can use numerical solutions to make inferences from the posterior distribution.
Regardless of the amount of information we have or how confident we are in that information, the prior distribution should at least take into account the domain (set of possible values) of the parameter. For example, if we want to estimate a proportion, the prior distribution for the parameter should range between 0 and 1.
Prior distributions could also be classified as conjugate priors. A conjugate prior is a prior that, when combined with the likelihood, results in a posterior distribution of the same form (parametric family) as the prior. An example of a conjugate prior is the beta prior for a binomial likelihood; the posterior distribution is also a beta distribution (see beta-binomial and Conjugate Prior). Conjugate priors are useful in the sense that the posterior distribution is of a known parametric form, thus the computations are less complicated and they can be interpreted as additional data. Even though conjugate priors have their advantages, their use is limited in real-world problems because they may not represent the prior knowledge appropriately for a specific situation, or no conjugate family exists for the real-world model. In situations where the form of the posterior distribution is not known (non-conjugacy), we can use numerical solutions to make inferences from the posterior distribution.
Posterior Distribution
It is possible to obtain an analytic solution when working with conjugate priors but, as the problem and the model get more complicated, these analytical solutions become less feasible. For most complex prior and likelihood combinations, the posterior distribution does not have a closed form, includes more than one parameter, or is the result of a hierarchical model. In such cases, a common solution is to use Markov Chain Monte Carlo (MCMC) algorithms such as Gibbs or Metropolis-Hastings sampler (see Gelman et al. 2014) to fit the model. MCMC is a computational approach that uses approximate distributions to sample values of θ, and at each iteration the samples are corrected so that the posterior distribution is better approximated in the next sampling. We expect that at some point the chains converge to the posterior distribution. In this process (shown in Figure 2), three Markov chains are used (red, black, and green), starting from different points and converging to a stationary distribution for the parameter σ. In this plot the x-axis is the MCMC iteration number, and on the y-axis is the parameter value drawn at each iteration. In a real analysis, the draws sampled before convergence (e.g., before iteration 4000 in Figure 2) are discarded and not used for the analysis; this is known as burn-in period.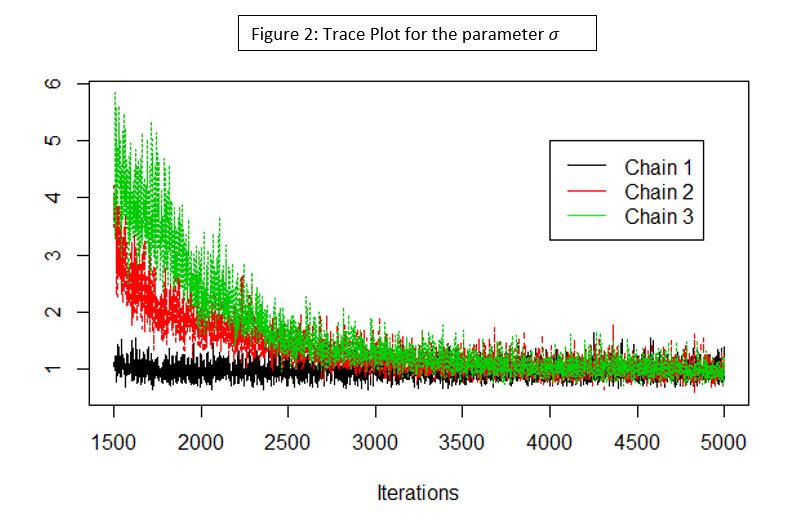 To ensure adequate coverage of parameter space and assess convergence, it is recommended that we run more than one chain with different initial values (a common rule of thumb is to use three chains). Note that the initial values selected for each chain might affect the convergence speed. That is, if the model is complicated and the initial values are extremely far away from each other, we might need to run thousands of iterations before model convergence. MCMC algorithms are available in software such as JAGS and OpenBUGS, and we can also write code to perform this process using R.
To ensure adequate coverage of parameter space and assess convergence, it is recommended that we run more than one chain with different initial values (a common rule of thumb is to use three chains). Note that the initial values selected for each chain might affect the convergence speed. That is, if the model is complicated and the initial values are extremely far away from each other, we might need to run thousands of iterations before model convergence. MCMC algorithms are available in software such as JAGS and OpenBUGS, and we can also write code to perform this process using R.
Model Diagnostics
Bayesian methods have techniques similar to those in frequentist analysis for checking the model fit. Once the model is fitted, a convergence assessment, as well as a check for model performance, must be implemented. Trace plots and Gelman and Rubin diagnostics are helpful in assessing convergence. The top image in Figure 3 shows a trace plot in which we can see that the three chains (red, green, and black) look like they are moving within a range and mixing well, which is a sign of convergence. Conversely, the bottom image on Figure 3 shows an example of chains that do not converge. We will also want to obtain a small MCMC standard error (MCMC standard error less than 5% of the posterior standard deviation of the parameter). If we do not obtain this initially, the most common way to achieve it is to run longer chains.
 If convergence is met, we can use Bayesian p-values (\(p_B\)) to compare the test statistics from replicated data (\(Y^{rep}\)) to the ones from the observed data (Y) . The Bayesian p-value can then be defined as follows:
[latex]p_B=Pr(T(Y^{rep},θ)>T(Y,θ)│Y) [/latex]
Contrary to the frequentist statistics, we do not want to obtain a small p-value but rather a Bayesian p-value as close to 0.5 as possible since that will mean that there is no disagreement between the replicated and the observed data. Bayesian p-values are used in posterior predictive checking as a way to assess whether the model is consistent with the data.
A sensitivity analysis should be performed to see how robust the model is to the specification of the prior. That is, how do the results change if we change the prior? If the end conclusions do not change, then we say that the model is robust to the selection of the prior. The specification of the prior has been criticized for being subjective since the analyst is the one who decides which prior to use in the analysis. A discussion of this limitation, as well as the advantages of Bayesian analysis, can be found in the Advantages and Disadvantages of Bayesian analysis section.
For more information, see:
Gelman, A., Carlin, J. B., Stern, H. S., Dunson, D. B., Vehtari, A., & Rubin, D. B. (2014). Bayesian data analysis (Vol. 2). Boca Raton, FL: CRC press.
Kruschke, J. (2014). Doing Bayesian data analysis: A tutorial with R, JAGS, and Stan. Academic Press.
If convergence is met, we can use Bayesian p-values (\(p_B\)) to compare the test statistics from replicated data (\(Y^{rep}\)) to the ones from the observed data (Y) . The Bayesian p-value can then be defined as follows:
[latex]p_B=Pr(T(Y^{rep},θ)>T(Y,θ)│Y) [/latex]
Contrary to the frequentist statistics, we do not want to obtain a small p-value but rather a Bayesian p-value as close to 0.5 as possible since that will mean that there is no disagreement between the replicated and the observed data. Bayesian p-values are used in posterior predictive checking as a way to assess whether the model is consistent with the data.
A sensitivity analysis should be performed to see how robust the model is to the specification of the prior. That is, how do the results change if we change the prior? If the end conclusions do not change, then we say that the model is robust to the selection of the prior. The specification of the prior has been criticized for being subjective since the analyst is the one who decides which prior to use in the analysis. A discussion of this limitation, as well as the advantages of Bayesian analysis, can be found in the Advantages and Disadvantages of Bayesian analysis section.
For more information, see:
Gelman, A., Carlin, J. B., Stern, H. S., Dunson, D. B., Vehtari, A., & Rubin, D. B. (2014). Bayesian data analysis (Vol. 2). Boca Raton, FL: CRC press.
Kruschke, J. (2014). Doing Bayesian data analysis: A tutorial with R, JAGS, and Stan. Academic Press.


